Francastel group
DNA methylation and non-coding RNA in health and disease

group by Zoom
The 4 members of the Francastel team: Claire Francastel, DR2 INSERM; Florent Hubé, CRCN CNRS; Guillaume Velasco, MCU Université Paris Cité; Katia Boyarchuk, Engineer CNRS.
© EDC
The overall interest of our team is on the multiplicity and intricacy of the regulatory networks that control expression and maintenance of mammalian genomes, from local transcriptional and epigenetic control to global genome organization within the nuclear space. With the genomes of complex eukaryotes being pervasively transcribed but mostly made up of large amounts of sequences that do not carry information to make proteins, a challenging task is to document this huge non-coding transcriptional output and understand its functional significance in regulatory networks that govern normal cell fate. We are particularly focused on understanding whether and how perturbations to this non-coding production is a driving force is disease, and which are the mechanisms and factors that maintain their normal biogenesis and hence, genome and cell integrity. Over the past years, we focused on inherently non-coding genomic regions that represent a vast fraction of mammalian genomes, i.e. repeated sequences and introns, as paradigms to address these questions and as unconventional targets of (post)transcriptional and epigenetic perturbations that underpin human diseases.
The strategies of our team are both basic and patient-oriented, based on (i) the development of mouse models, (ii) the establishment of cohorts of patients in collaboration with physicians, (iii) a combination of complementary expertise in the study of gene expression, DNA methylation, non-coding RNA and their associated complexes, RNA splicing and processing and nuclear architecture, and (iv) capitalize on unique cellular models that we developed, including mouse models, ES cells and cells from patients, and the generation of high throughput data. Technical approaches include classical molecular biology, high throughput techniques for epigenome and transcriptome analysis, cellular imaging and in situ assays in the mouse.
Our main goals are to:
– document the non-coding transcriptional output from these regions in normal and physiopathological situations
– characterize the biogenesis of the produced transcripts, their processing, epigenetic regulation, sub-cellular localization and associated complexes
– understand if and how their deregulation represents a driving force in disease
– provide an integrated view of pan-genomic epigenetic, transcriptional and splicing defects in the context of pathological defects of the DNA methylation machinery.
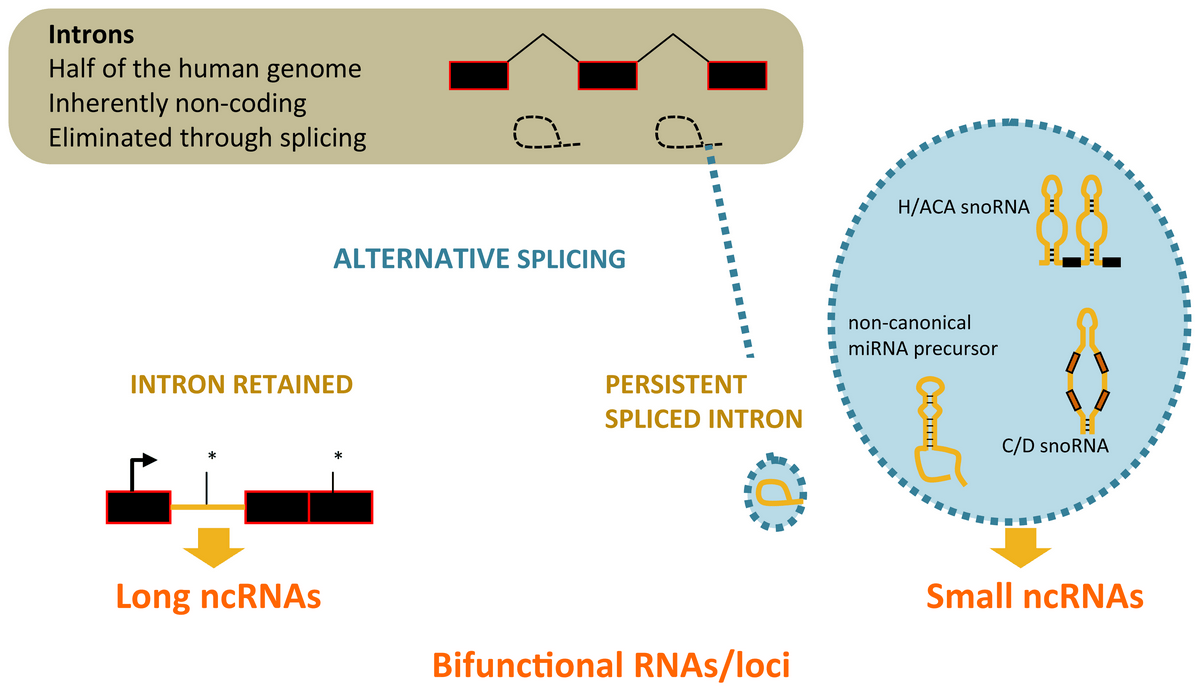
Alternative splicing of introns and the diversification of transcriptional output
A paradigm to test the functionality of non-protein-coding genomic regions
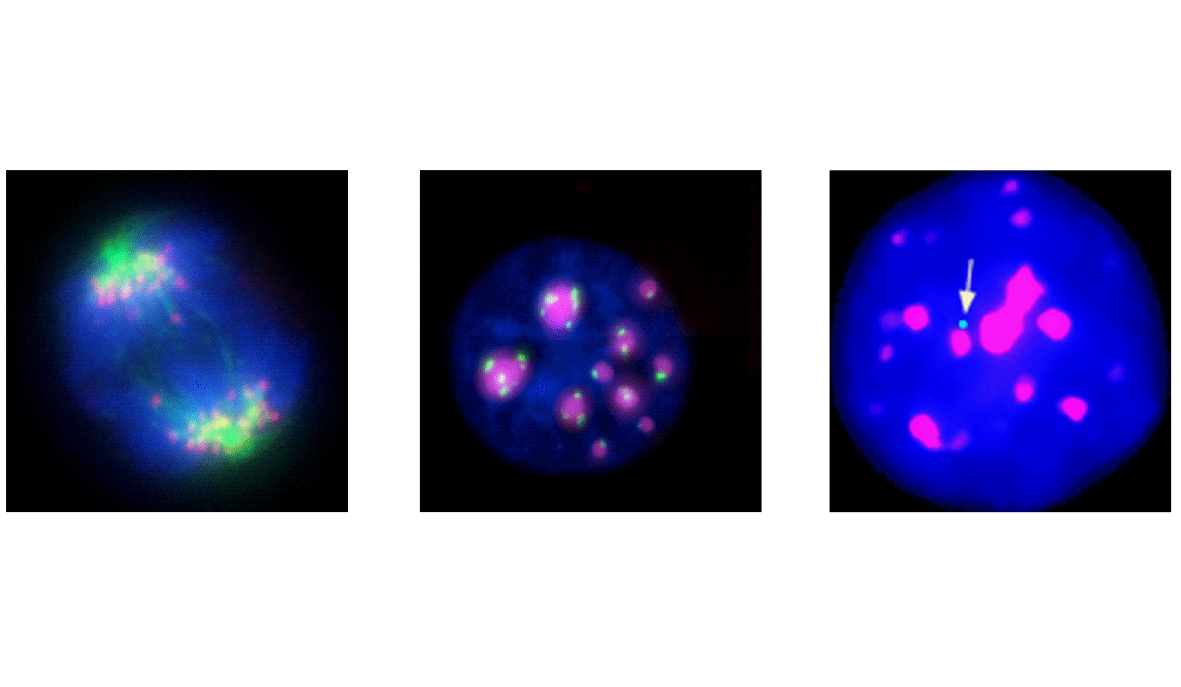
Centromeric repeats transcription
A paradigm to link transcription of DNA repeats to global molecular and cellular effects
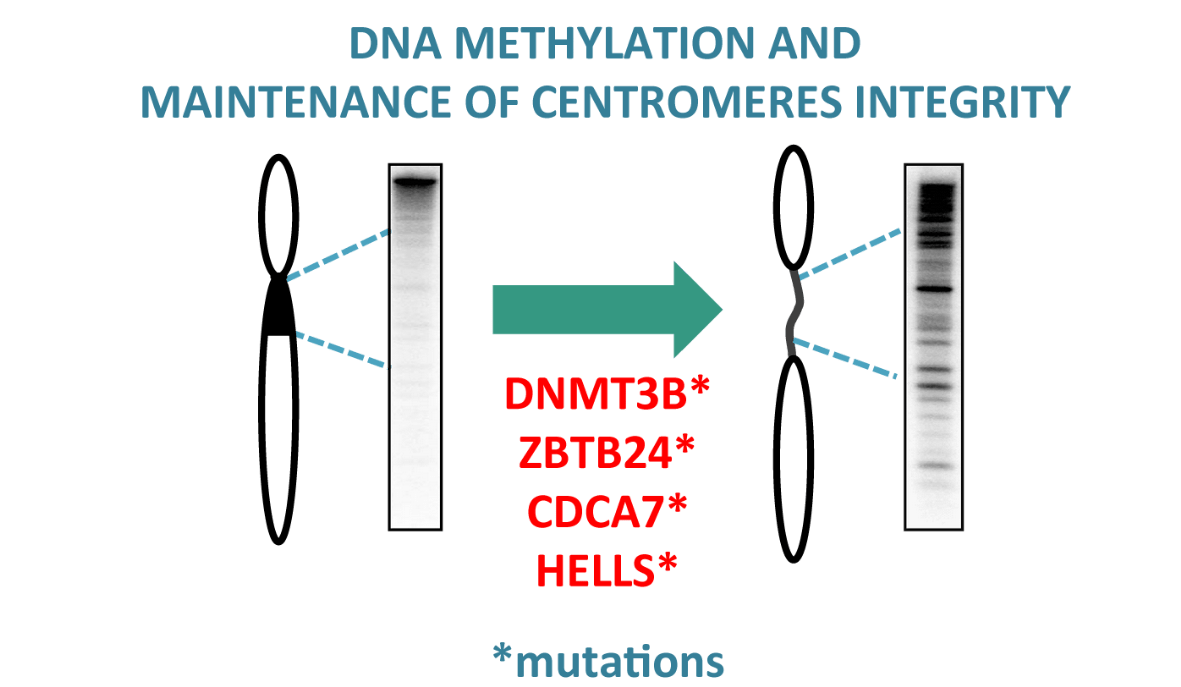
DNA methylation and genome maintenance
When studying a rare disease sheds new light on the field of DNA methylation
Publications
Contact
Membres
Read more

Seminar Anne-Sophie Lebre– May 27, 2026
Pr. Anne-Sophie Lebre CHU de Reims - Hôpital Robert Debré, France Invited by Pr. Bertrand Cosson "Brain (epi)genome study in CNS rare disorders" The seminar will take place in the Institut Jacques Monod seminar room (RB-18B). Bâtiment Buffon, 15 rue Hélène Brion,...

Seminar Hiroshi Kimura– May 20, 2026
Dr. Hiroshi Kimura Institute of Quantitative Biosciences, University of Tokyo, Japan Invited by Dr. Pierre-Antoine Defossez "Dynamics of chromatin modification and transcription in vivo." The seminar is Zoom-only On Wednesday, May 20that 11:00 am....

Bing Zhu Seminar – May 6, 2026
Dr. Bing Zhu Institute of Biophysics, Chinese Academy of Sciences, Beijing, China Invited by Dr. Pierre-Antoine Defossez "Epigenetics: remember the past & prepare for the future." The seminar is Zoom-only On Wednesday, May 6that 11:00 am....

Harm H. Kampinga Seminar – April 15, 2026
Dr. Harm H. Kampinga Dept. Biomedical Sciences, UMCG/RuG, Groningen, The Netherlands Invited by Dr. Valérie Mezger "Collapse of protein homeostasis as driver of age-related protein aggregation diseases" The seminar will take place in the Institut Jacques Monod seminar...
