EpiG : Functional epigenomics
The Functional Epigenomics platform’s objectives are to provide equipment and to train staff in its use.
It offers services for epigenome analysis and participates in the development of new techniques according to the evolution of the field.
The Functional Epigenomics platform is open to the academic and industrial scientific community.
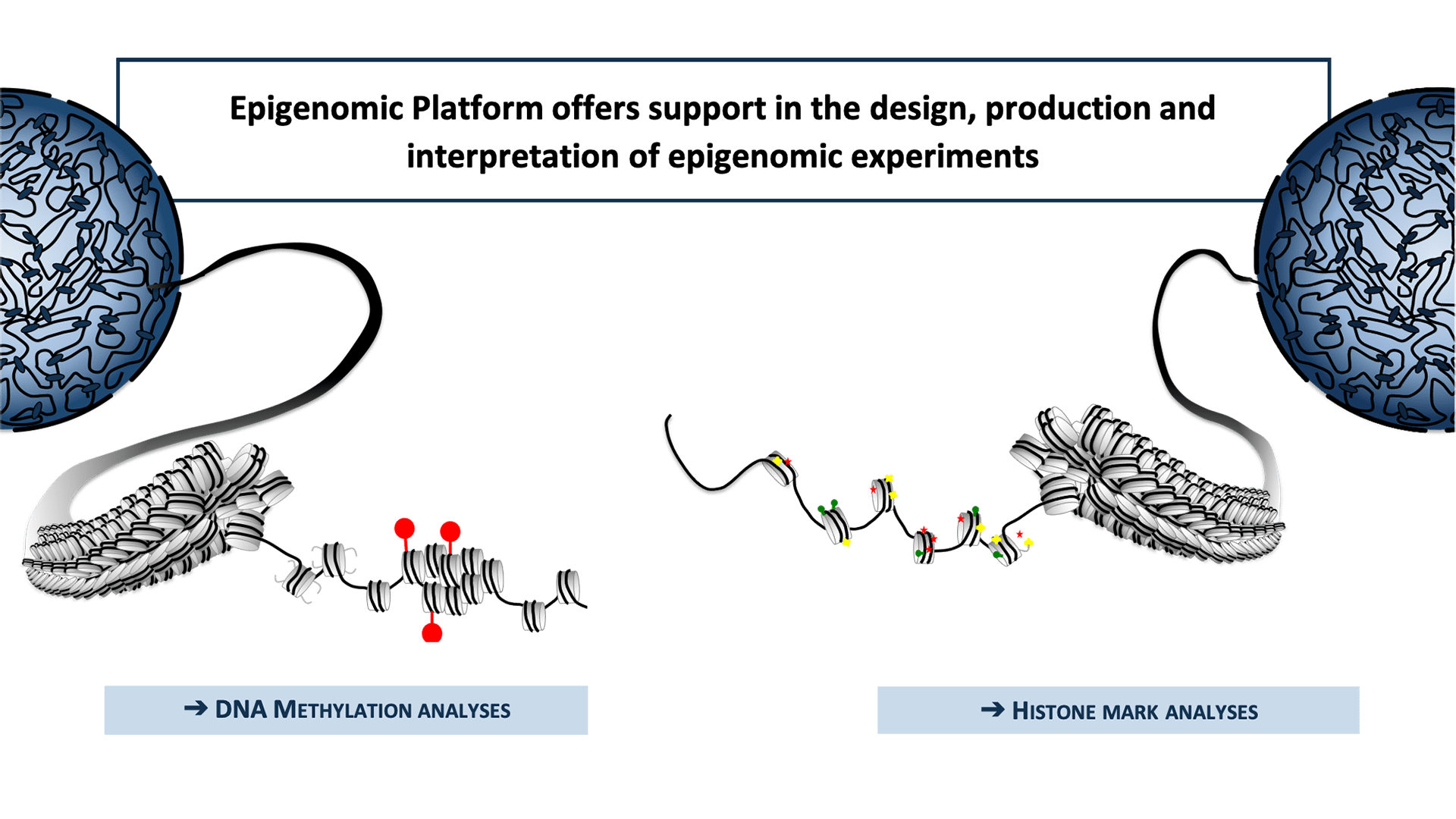

Scheme of platform’s expertise
Epigenomic Platform offers support in the design, production and interpretation of epigenomic experiments.
© Laure Ferry
Expertise
The epigenomic platform offers expertise in analysis of epigenetic marks. It evaluates the feasibility of submitted projects and proposes the most suitable method for their completion.
DNA Methylation
- LUMA (LUminometric Methylation Assay)
- MeDIP ou hMeDIP (Methylated or hydroxyMethylated Immunoprecipitation Assay)
- Bisulfite pyrosequencing
Histones Marks
- Cut & Run
- ChIP
Submit a project
You can contact us for any questions or to discuss your project together.
You must fill in the project request form and send it to the following address: parisepigenetics.pf@gmail.com
Developments
The Functional Epigenomic platform is involved in the development of new techniques for epigenome analysis:
- Cut & Run automatisation
- Cut & Tag
- Enzymatic Methyl-seq ( EM-seq)
- Nanopore
Epigenetic analyses available
Services for DNA methylation and histone marks analyses
Equipment
Equipment available on the Epigenomic platform
Members
Publications
- Planques A, Pierre K, Ferry L, Grunau C, Gazave E, Vervoort M. DNA methylation atlas and machinery in the developing and regenerating annelid Platynereis dumerilii. BMC Biology, Vol 19, Iss 1, Pp 1-26, 2021. DOI: 10.1186/s12915-021-01074-5
- Marchal C, de Dieuleveult M, Saint-Ruf C, Guinot N, Ferry L, Olalla Saad ST, Lazarini M, Defossez PA, Miotto B. Depletion of ZBTB38 potentiates the effects of DNA demethylating agents in cancer cells via CDKN1C mRNA up-regulation. Oncogenesis; Oct 2018, Vol. 7 Issue 10, p1-18, 18p DOI: 10.1038/s41389-018-0092-0
- Velasco G, Grillo G, Touleimat N, Ferry L, Ivkovic I, Ribierre F, Deleuze JF, Chantalat S, Picard C, Francastel C. Comparative methylome analysis of ICF patients identifies heterochromatin loci that require ZBTB24, CDCA7 and HELLS for their methylated state. Human Molecular Genetics, 2018 Jul 15;27(14):2409-2424. DOI: 10.1093/hmg/ddy130.
- Ferry L, Fournier A, Tsusaka T, Adelmant G, Shimazu T, Matano S, Kirsh O, Amouroux R, Dohmae N, Suzuki T, Filion GJ, Deng W, de Dieuleveult M, Fritsch L, Kudithipudi S, Jeltsch A, Leonhardt H, Hajkova P, Marto JA, Arita K, Shinkai Y, Defossez PA. Methylation of DNA Ligase 1 by G9a/GLP Recruits UHRF1 to Replicating DNA and Regulates DNA Methylation. Molecular Cell. August 17, 2017, Vol. 67 Issue 4, p550. DOI: 10.1016/j.molcel.2017.07.012
- Kebir O, Chaumette B, Rivollier F, Miozzo F, Lemieux Perreault LP, Barhdadi A, Provost S, Plaze M, Bourgin J, the ICAAR team, Gaillard R, Mezger V, Dubé M-P , Krebs M-O. Methylomic changes during conversion to psychosis. Molecular Psychiatry (2017) 22, 512-518. DOI: 10.1038/mp.2016.53.
Contact
You can contact us at: parisepigenetics.pf@gmail.com
Functional Epigenomic Platform
35 rue Hélène Brion
Bâtiment Lamarck B, RB77-81
75013 PARIS
+33(0)1 57 27 89 26
Read more

Seminar Anne-Sophie Lebre– May 27, 2026
Pr. Anne-Sophie Lebre CHU de Reims - Hôpital Robert Debré, France Invited by Pr. Bertrand Cosson "Brain (epi)genome study in CNS rare disorders" The seminar will take place in the Institut Jacques Monod seminar room (RB-18B). Bâtiment Buffon, 15 rue Hélène Brion,...

Seminar Hiroshi Kimura– May 20, 2026
Dr. Hiroshi Kimura Institute of Quantitative Biosciences, University of Tokyo, Japan Invited by Dr. Pierre-Antoine Defossez "Dynamics of chromatin modification and transcription in vivo." The seminar is Zoom-only On Wednesday, May 20that 11:00 am....

Bing Zhu Seminar – May 6, 2026
Dr. Bing Zhu Institute of Biophysics, Chinese Academy of Sciences, Beijing, China Invited by Dr. Pierre-Antoine Defossez "Epigenetics: remember the past & prepare for the future." The seminar is Zoom-only On Wednesday, May 6that 11:00 am....

Harm H. Kampinga Seminar – April 15, 2026
Dr. Harm H. Kampinga Dept. Biomedical Sciences, UMCG/RuG, Groningen, The Netherlands Invited by Dr. Valérie Mezger "Collapse of protein homeostasis as driver of age-related protein aggregation diseases" The seminar will take place in the Institut Jacques Monod seminar...
