BiBs : Bioinformatics and Biostatistics
The Epigenetics and Cell Fate lab BiBs facility provides a broad range of services related to bioinformatics.


Picture from Gerd Altmann, source: Pixabay.
Presentation
The BiBs platform is part of and actively contributes to the development of the iPOP-UP platform which federates and promotes bioinformatics at the level of the Université Paris Cité.
The facility includes two permanent staff and regularly welcomes trainees or apprentices.
Expertise
Our expertise is about the analysis of high-throughput sequencing data (Illumina and Oxford Nanopore). We can guide you on analyzing different types of omics data:
- Transcriptome (RNA-seq)
- Methylome (bisulphite sequencing, or direct long-read detection by nanopore)
- Genomic localization of transcription factors or histone marks (ChIP-seq or CUT&RUN)
- Chromatin accessibility (ATAC-seq)
- Chromosome conformation (Hi-C)
In addition to ‘bulk’ analyses, we are strengthening our skills in the analysis of ‘single-cell’ data, in particular thanks to the iPOP-UP network.
We are also experts in the integration of different bioinformatics tools within workflows (Snakemake), which makes analyzes reproducible and accessible to a wide audience of researchers. Our workflows are compatible with the high performance computing clusters of the Université Paris Cité (iPOP-UP, hosted by RPBS) and the French Institute of Bioinformatics.
Finally, we have expertise in data management (drafting of data management plans, FAIR principles).
Services
-
Experimental design: advice on the choice of sequencing technique, the necessary depth, the controls, the number of biological replicates…
-
Development of high throughput sequencing data analysis workflows and support in their use.
-
Support in the use of the iPOP-UP high-performance computing cluster and organization of dedicated training.
-
Maintenance and promotion of the bioinformatics tools developed by members of the laboratory.
-
On-demand training on the use of bioinformatics tools to develop user autonomy.
-
Proofreading of articles or funding requests concerning projects using NGS analyses.
-
Assistance in the recruitment of interns, doctoral students, post-doctoral fellow or engineers in charge of omics data analysis.
-
Best practice recommendations for data management and analysis based on the FAIR principles.
Members
Magali Hennion
Head, IR CNRS
Olivier Kirsh
Assistant professor, Université Paris Cité
Mélina Farshchi
Trainee, master M2-BI
Resources
Website of the facility
All tutorials and related documentation written by BiBs are available on https://parisepigenetics.github.io/bibs/.
GitHub
All codes developed by EDC lab members are available on https://github.com/parisepigenetics.
Contact
BiBs
Lamarck B building, room 366b
35 rue Hélène Brion, 75013 Paris
e-mail: bibs@parisepigenetics.com
Steering committee
Read more

Congratulations to the Defossez group for their review published in Médecine/Sciences
Congratulations to Matthieu, Raady and Pierre-antoine for their review "Demethylating DNA against cancer : towards new epigenetic therapeutic strategies without DNA damage" published in Médecine/Sciences in february 2026! Read here
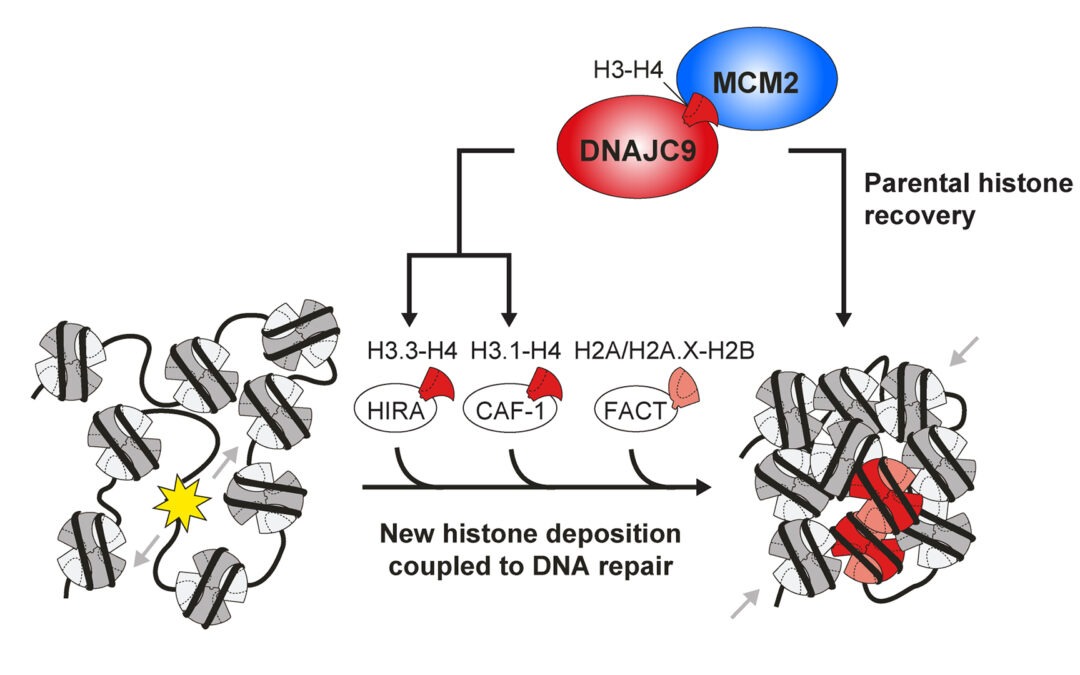
New research article on histone chaperones with central functions in chromatin repair
In this paper, we identify new players in chromatin repair through a novel approach for the proteomic profiling of DNA repair patches in human cells. Thus, we expand the histone chaperone network involved in repair-coupled chromatin restoration and highlight a...

Welcome to Léa
Léa joins the team as a research assistant. After completing a master's degree in virology, she worked in Strasbourg on grapevine viruses, then on characterizing mRNA degradation in plants at the Institute of Plant Molecular Biology (IBMP). In the Polo team, Léa will...

Sophie Polo receives an Impulscience® grant from the Fondation Bettencourt Schueller
Sophie Polo has been awarded an Impulscience® grant to fund a research project on the establishment and maintenance of the inactive X chromosome in response to DNA breaks. This is wonderful news for the lab ! We thank the Fondation Bettencourt Schueller for their...
